|
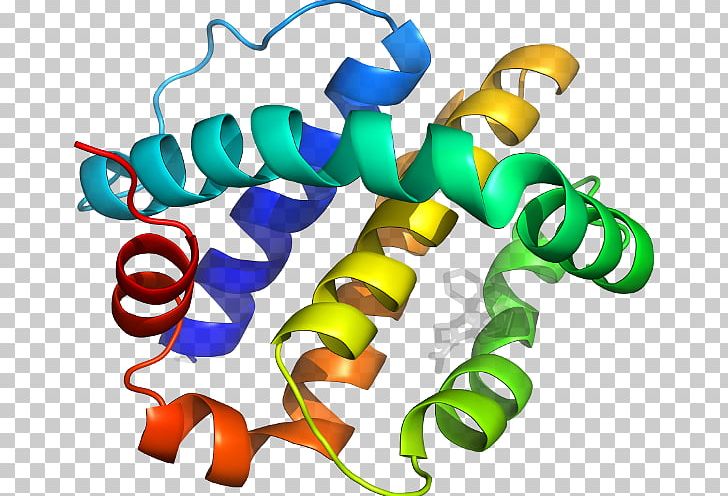
For each protein, click Download Files and select PDB Format. Throughout this tutorial, the program may prompt you to download additional protein database information, which you should accept. Getting startedįor this tutorial, first download VMD. We will update this module when the software is fixed. Note: As of the current time, the STAMP alignment step used by Multiseq is not working for most users, especially Windows users. For more details on the algorithm used by STAMP, click here. If the structures do not have common structures, then STAMP will fail. Much like the Kabsch algorithm considered in part 1 of the module, STAMP minimizes the distance between alpha carbons of the aligned residues for each protein or molecule by applying rotations and translations. Multiseq aligns two protein structures using a tool called Structural Alignment of Multiple Proteins (STAMP). By locating regions with low Qres, we can hopefully identify regions of structural differences between the two RBDs. In this tutorial, we will get started with VMD and then calculate Qres between the SARS-CoV-2 RBD (PDB entry: 6vw1) and SARS-CoV RBD (PDB entry: 2ajf) using the VMD plugin Multiseq.
0 Comments
Leave a Reply. |




 RSS Feed
RSS Feed
